Last night I gave a presentation at the Drupal WA Meetup on a recent project we’ve collaborated on with the Western Australian Museum. The aim of the project was to make up to date species identification information available to field workers. I’d like to give a little background about the project and some details as to how we went about integrating the Museum’s collections databases with a Drupal frontend to give environmental consultants a dynamic identification aid.
Collections and Databases
Over the years we have worked with the Western Australian Museum (WAM) on a number of projects including database and GIS support. They have been a great client to work with and we have learned a lot from eachother. Over the past few months I’ve been spending a couple of days a week as an embedded developer at the Museum working on a project to ‘Integrate the collections databases with the Museum’s online catalogues’. After a few days of discussions with stakeholders and getting myself setup with the Museum’s development framework of Drupal on a fairly beefy LAMP server I set out on the project that became affectionately known as ‘Waminals’. Each collection at the Museum has it’s collections data housed in a separate database that has been tailored to that particular collection’s needs. Here are a few details about the previous workflow and what we were trying to attain.
Previous Workflow
The previous workflow was:
- Consultant lodges specimens, collecting data and identifications with the WAM
- WAM curator then receives specimens
- Checks identifications
- Checks taxonomy
- Assigns ‘putative name’ while taxa are still being described
- Accessions Specimens
- Adds the specimen to the museum’s catalogue (database) and stores it in the appropriate location and medium.
- Gives consultant back a list of accession numbers, collecting information and the correct identification
- Often these names are not fully published names but rather field names that will eventually get replaced by validly published scientific names
- The consultant is then ignorant of any subsequent changes to the data
Waminals Workflow
The Waminals project aimed to enhance this workflow by creating factsheets for taxa that are commonly thought of as new
- Created by the ecological consultants on museum infrastructure
- Checked by museum curators and approved for ‘publication’
- Consultant will add
- descriptive couplets (attributes) of character name and character state
- Multiple images of specimens which have been registered in the WAM collections
- Registration numbers for the specimens in the photographs
- Name details that the consultant has are entered at the same time
- Factsheet then checked by WAM staff, and when approved it will be published.
- The System will then periodically
- Take the registration numbers from the factsheet
- Query collection databases for names associated with those registration keys
- Update the name details on the factsheet to match the current name used in the CMIS.
- These fields should then be locked from further update by the users – updates now come from the database.
- ‘Putative names’ will eventually be replaced by the correct name when published.
- Retrieve other specimens that are associated with that name
- Pull in coordinates & locality information for these specimens
- Display them in an online map
- The system should only expose data to authorised users due to the commercial and intellectual property sensitivities of the data.
Data Transformations
 Due to the heterogeneous database backends at the museum we could not simply expose the collections data to Drupal without an intermediate transformation step. Luckily there are plenty of people out there who have been in the business of transformations a long time and have created great tools such as Google Refine (see my previous blog post on using that) and Pentaho Data Integration. The Museum is currently using Pentaho to mobilise some of their collections data to the Atlas of Living Australia, and it provides the facility to run the transformations server side via a shell script, so the decision to use it was an easy one to make.
Due to the heterogeneous database backends at the museum we could not simply expose the collections data to Drupal without an intermediate transformation step. Luckily there are plenty of people out there who have been in the business of transformations a long time and have created great tools such as Google Refine (see my previous blog post on using that) and Pentaho Data Integration. The Museum is currently using Pentaho to mobilise some of their collections data to the Atlas of Living Australia, and it provides the facility to run the transformations server side via a shell script, so the decision to use it was an easy one to make.
Data exchange format
 So now we had a tool to do the data transformations for us we then needed to settle on a standard exchange format for the specimen data. Again we didn’t have to look much further than TDWG and their DarwinCore standard as it now supports the lowest common denominator of data exchange formats – CSV.
So now we had a tool to do the data transformations for us we then needed to settle on a standard exchange format for the specimen data. Again we didn’t have to look much further than TDWG and their DarwinCore standard as it now supports the lowest common denominator of data exchange formats – CSV.
<rant>CSV stands for ‘Character Separated Values’ not ‘Comma Separated Values’.</rant>
Data Loading
There are a number of ways to expose data to Drupal from quick and dirty hacks to elaborate modules with complex data loading regimes, and we evaluated the Feeds and the Migrate modules. Feeds has a nice UI and you can map your source data to Drupal fields through point and shoot functionality, but it missed a few features that we were looking for. Migrate on the other hand has a fairly simple UI and the mapping of data to Drupal entities is done through extending the base Migration class in PHP code. This meant that we could define fairly complex transformations during data loading. Additionally the Feeds module appears to be suffering from a little bit rot lately with the bug number creeping ever higher.
Site Structure
Wherever possible we used Drupal modules that exposed their configurations to the excellent Features module which allows you to export and version control your configurations making deployment between staging and production servers much less of a headache than essentially ‘doing the same thing again on a different server’. Additionally all the code developed was pushed to the Museum’s Git repository to make collaboration easier.
Factsheets
The Museum’s Catalogues Site is where all the data had to end up. The guys at the Museum had already set up a content type to capture the factsheet data in using Drupal’s great Field UI. Species descriptions often take a similar form, but depending on the group you are working with the names of characters and states can vary. The WAM developers came up with a neat solution to this by using hook_field_info() to define a field type that gives you two fields per value, one for the character name and another for the character state. The character name autocompletes from pre-existing values helping to normalise number of characters entered. The nice thing about using Drupal’s field system is that these fields get made available to the Views module so you can start building complex queries on them.
Consultants from collaborating ecological consultancies then filled in the species descriptions and uploaded images throughout the process of the project.
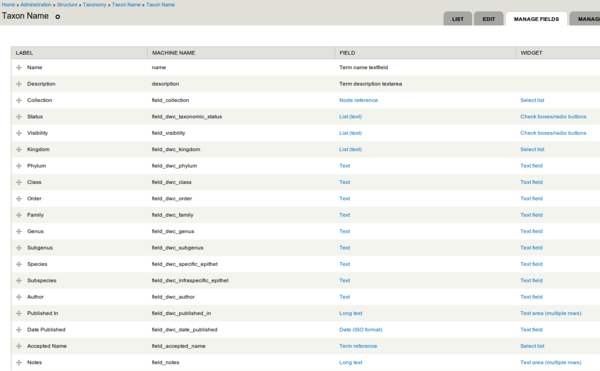
Taxonomy
The names from the specimens get stored in a Drupal Taxonomy containing additional fields inspired by DarwinCore Taxon level terms.
Occurrences
The specimens were loaded into a Drupal content type that was modelled around the Occurence, Event and Location level terms with references to the Taxon vocabulary within the site.

Geospatial Data
 Geospatial data in Drupal has had a fairly complex history and the number of spatial modules is quite overwhelming. Luckily the Geofield module has brought some kind of consistency, embracing the above mentioned Field UI. This means that you can add any number of spatial fields to a Drupal entity, each field with their own settings and input widgets, so you could for example have a lat/lon pair of fields for a specific point and then have a separate draw on web map to define the polygon that you’re interested in (think capital city in a country, or location of specimen within a collecting area). The geofield module uses the geoPHP library (https://github.com/phayes/geoPHP, http://drupal.org/project/geophp) which in turn uses the PHP bindings for the GEOS (Geometry Engine Open Source) if you have them enabled. This means you can perform spatial transformations from within Drupal.
Geospatial data in Drupal has had a fairly complex history and the number of spatial modules is quite overwhelming. Luckily the Geofield module has brought some kind of consistency, embracing the above mentioned Field UI. This means that you can add any number of spatial fields to a Drupal entity, each field with their own settings and input widgets, so you could for example have a lat/lon pair of fields for a specific point and then have a separate draw on web map to define the polygon that you’re interested in (think capital city in a country, or location of specimen within a collecting area). The geofield module uses the geoPHP library (https://github.com/phayes/geoPHP, http://drupal.org/project/geophp) which in turn uses the PHP bindings for the GEOS (Geometry Engine Open Source) if you have them enabled. This means you can perform spatial transformations from within Drupal.
Online Maps
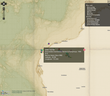 Again there were a few options with respect to how we were going to present the data from the Waminals project, but the OpenLayers project is the one that we have the most experience with and it’s Drupal implementation is fully featured and there are a good number of layer types available for Drupal, either build in or through modules such as the MapBox module.
Again there were a few options with respect to how we were going to present the data from the Waminals project, but the OpenLayers project is the one that we have the most experience with and it’s Drupal implementation is fully featured and there are a good number of layer types available for Drupal, either build in or through modules such as the MapBox module.
As the data we were presenting was not going to be available to freely registered users on the site due to commercial and intellectual property sensitivity we could not use Google Maps. There are however great datasets with decent imagery for the scale of maps that we were wanting to present including the NASA Blue Marble imagery.
Access Control
As mentioned above by default the data must not be exposed to unregistered users. This was achieved via the Content Access module which means that if the system gets extended and made publicly available it is simply a matter of specifying the access level of individual occurrences / factsheets and the system will expose them if necessary.
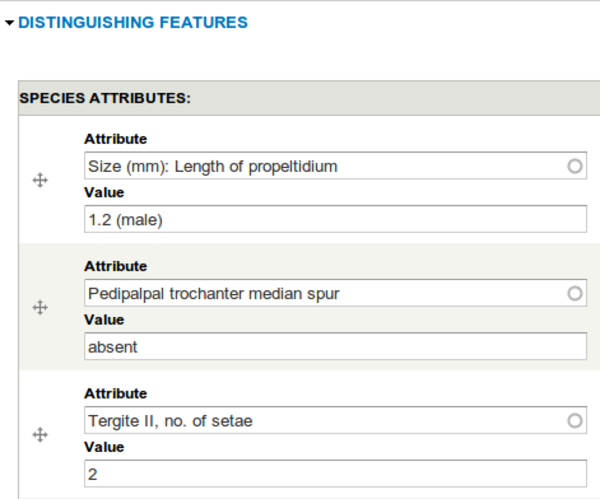
Sample Factsheet
Factsheet Browsing
We needed a way of browsing between factsheets and finding taxa with similar characteristics and the Views module was the starting point for that. A tabular view looked like what we wanted but we wanted the fields down the left hand side and the individual nodes as a side scrolling table. Again we did a bit of research and the Views Hacks module provides exactly that functionality in the form of a views_flipped_table output style. Some jQuery and CSS magic from the Museum’s designers and we had a nice tablet friendly factsheet browser thingamajig. Plus some careful url crafting meant that it was possible to pass in the view’s input filters in as GET query parameters so drilling down to factsheets with attributes matching the specimen being observed is quite fast and intuitive.
These factsheets do not replace classical dichotomous keys or interactive keys but the data captured through the collaborative infrastructure of a Drupal website makes a great starting point for creating either of these identification aids should the need arise.
Conclusion
It was great to be working on site with the client as queries could be answered quickly and changes could be tested as soon as they were implemented. Drupal is a great web framework and has come on leaps and bounds since I first started using it six years ago. Additionally, and more interestingly for us as a company with a fairly strong interest in spatial technology, it is possible to create web maps using Drupal without too many hassles so watch this space.
Presentation
I have embedded the presentation that I gave below if you would like to take a look:
If you have any responses contact me via email or twitter or leave a message below.



Comments are closed.